
First, a substantial part (up to 20%) of short Okazaki fragments (~150-nt in average) is synthesized by Pol α which does not contain a proofreading function ( Conaway and Lehman, 1982 Bullock et al., 1991). Lagging strand maturation appears to be intrinsically at high risks of suffering DNA alterations for several reasons. Potential risks associated with lagging strand synthesis in eukaryotes The nicks are finally sealed by DNA ligase 1 to complete Okazaki fragment processing. 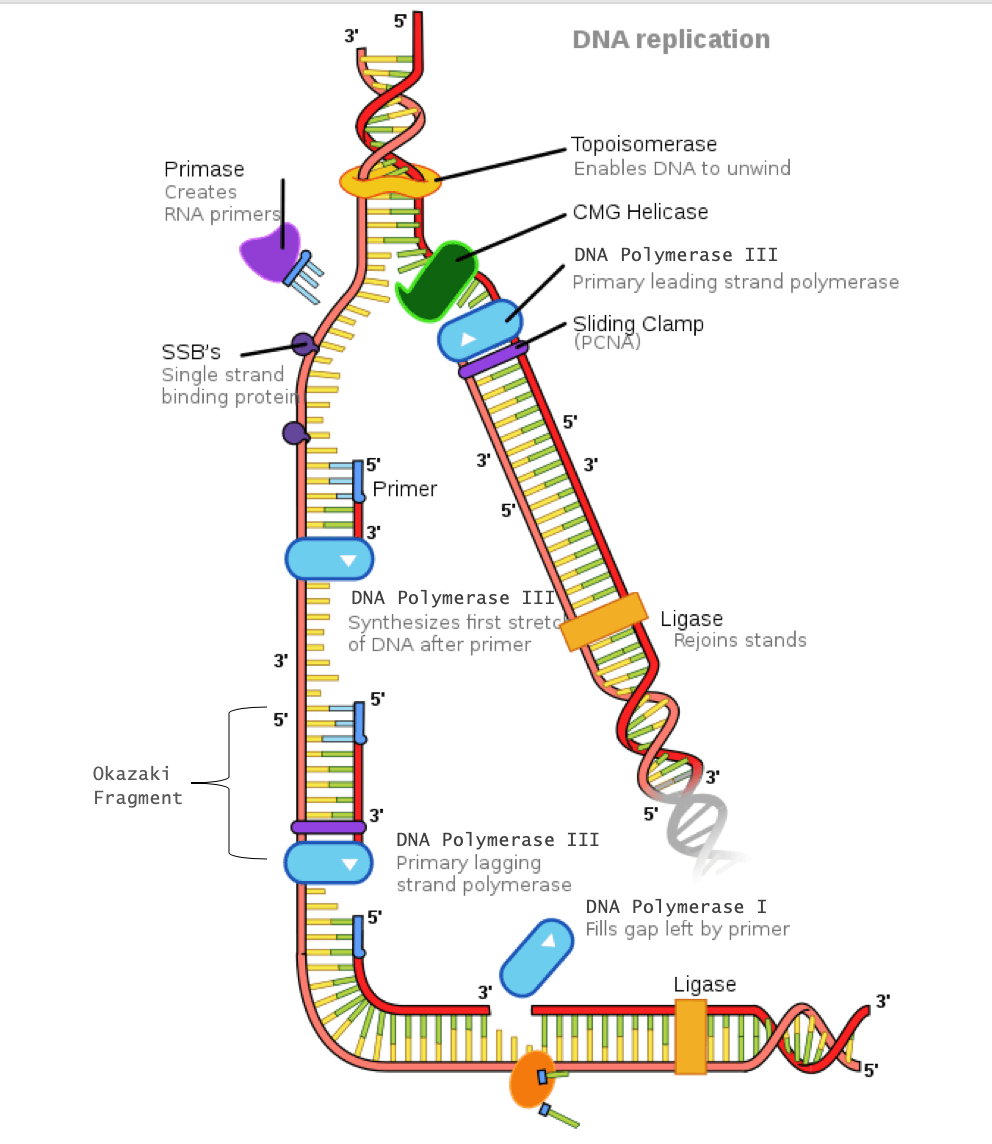
The flaps formed can be efficiently processed by the combined action of Flap endonuclease 1 (Fen1) and Dna2 to eventually create nicks. When Pol δ encounters a downstream Okazaki fragment, it displaces the 5’ end region of the Okazaki fragment, generating a single-stranded (ss) nucleic acid flap. The recognition of a primer RNA-DNA by the Replication-Factor C (RFC) complex leads to dissociation of Pol α-primase and loading of proliferating cell nuclear antigen (PCNA), resulting in recruitment of Pol δ to the primer-template junction, a process called ‘polymerase switching.’ Then the primer RNA-DNA is elongated by Pol δ. The synthesis of RNA portion (~ 10 to 15 ribonucleotides) and subsequent extension of short (~20 to 30 nucleotides, nt) DNA are coupled. Synthesis of an Okazaki fragment begins with a primer RNA-DNA made by polymerase (Pol) α-primase. Unlike leading strands, lagging strands are synthesized as discrete short DNA fragments, termed ‘Okazaki fragments’ which are later joined to form continuous duplex DNA. Leading strand synthesis, once initiated, occurs in a highly processive and continuous manner by a proofreading DNA polymerase. Activation of origins leads to the establishment of bidirectional replication forks for the DNA synthesis of leading and lagging strands. The assembly of replication initiation complex and its activation are well reviewed in many literatures ( Sclafani & Holzen, 2007 Remus & Diffley, 2009 Araki, 2010). 
The irreversible removal of initiation factors is a major mechanism to ensure DNA to be replicated once and only once per cell cycle. The elevated levels of Cdk activities lead to removal of some initiation proteins such as Cdc6 by proteolysis, allowing the pre-RC to be further activated for subsequent DNA synthesis. This complex is activated by S-phase cyclin dependent kinases (Cdks) when cells enter S phase. The prereplicative complex (pre-RC) is formed by loading the replicative helicase MCM complex onto the ORC-bound origins with the aid of Cdc6 and Cdt1. Briefly, the ORC-bound origins are sequentially activated and deactivated along the progression of cell cycle. In yeast and most likely higher eukaryotes, the origin-recognition complex (ORC) and several other initiation factors play a pivotal role in activation and regulation of replication origins. Each origin is activated at different time points in S phase, which takes place once and only once per cell cycle. The telomerase attaches to the end of the chromosome, and complementary bases to the RNA template are added on the end of the DNA strand.In eukaryotic cells, DNA replication starts at many origins in each chromosome during S phase of cell cycle. \)) helped in the understanding of how chromosome ends are maintained.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
